We apply a variety of computational tools and resources to help research teams solve key challenges in new drug R&D, such as biological activity, selectivity, and metabolic stability.
Structure-Based Drug Design
- Molecular docking and virtual screening
- Homology modeling
- De novo molecule generation and optimization
Ligand-Based Drug Design
- Pharmacophore modeling and virtual screening
- Molecular conformational analysis
- Quantitative structure-activity relationship (QSAR)
Calculation of Drug Metabolism and Physicochemical Properties
- Fast prediction of compound physicochemical properties
- In silico simulation of ligand or structure-based hERG and P450 pathways
Service Content 1: Protein 3D Structure Modeling – Homology Modeling or De Novo Modeling
Currently, there are three main methods to obtain protein structures: X-ray, NMR, and cryo-electron microscopy. If such experimental conditions are unavailable, computational simulation can resolve the 3D spatial structure of unknown proteins to support your research projects. Alternatively, we can calculate the 3D structure via homology modeling or de novo modeling, facilitating subsequent pharmacology and molecular biology studies.
Homology modeling (via Modeller software) uses known protein tertiary structures as templates to predict the 3D structure of a new protein with a similar amino acid sequence. For proteins with low sequence similarity, de novo modeling (via Rosetta software) can be applied to predict the 3D structure.
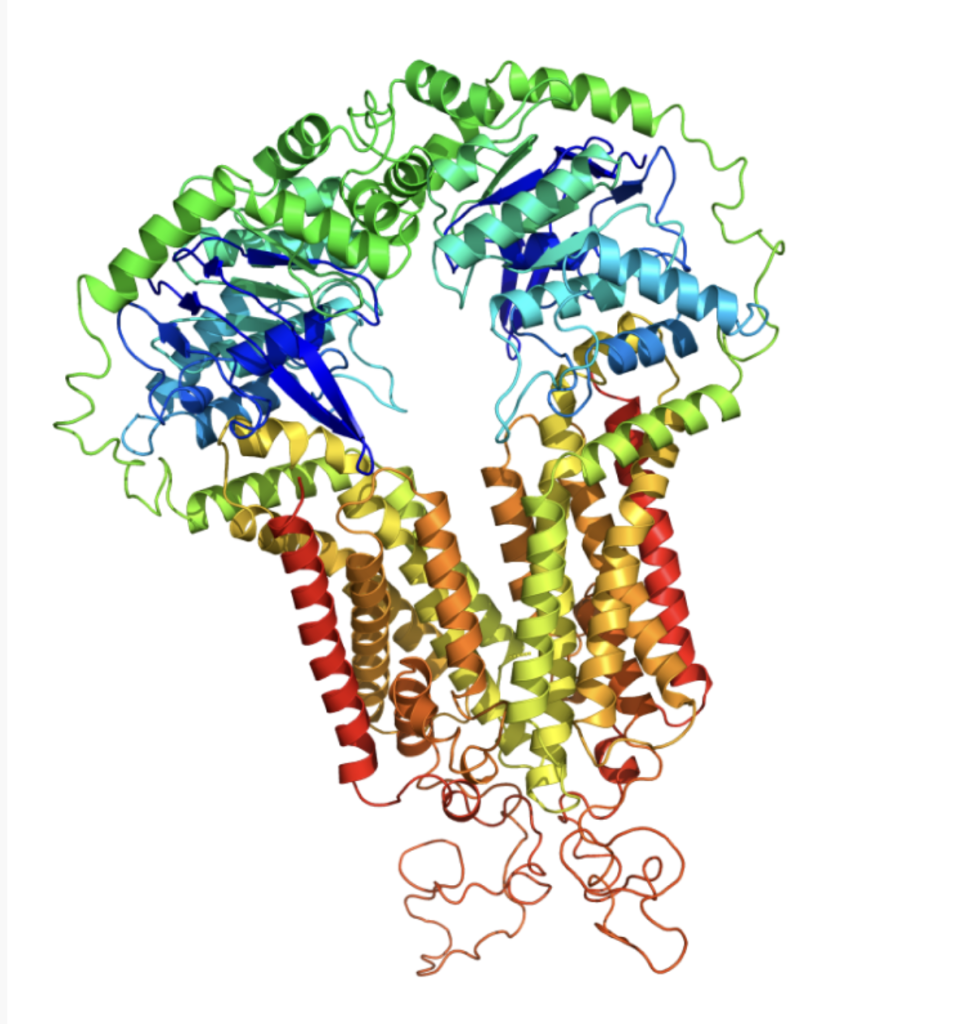
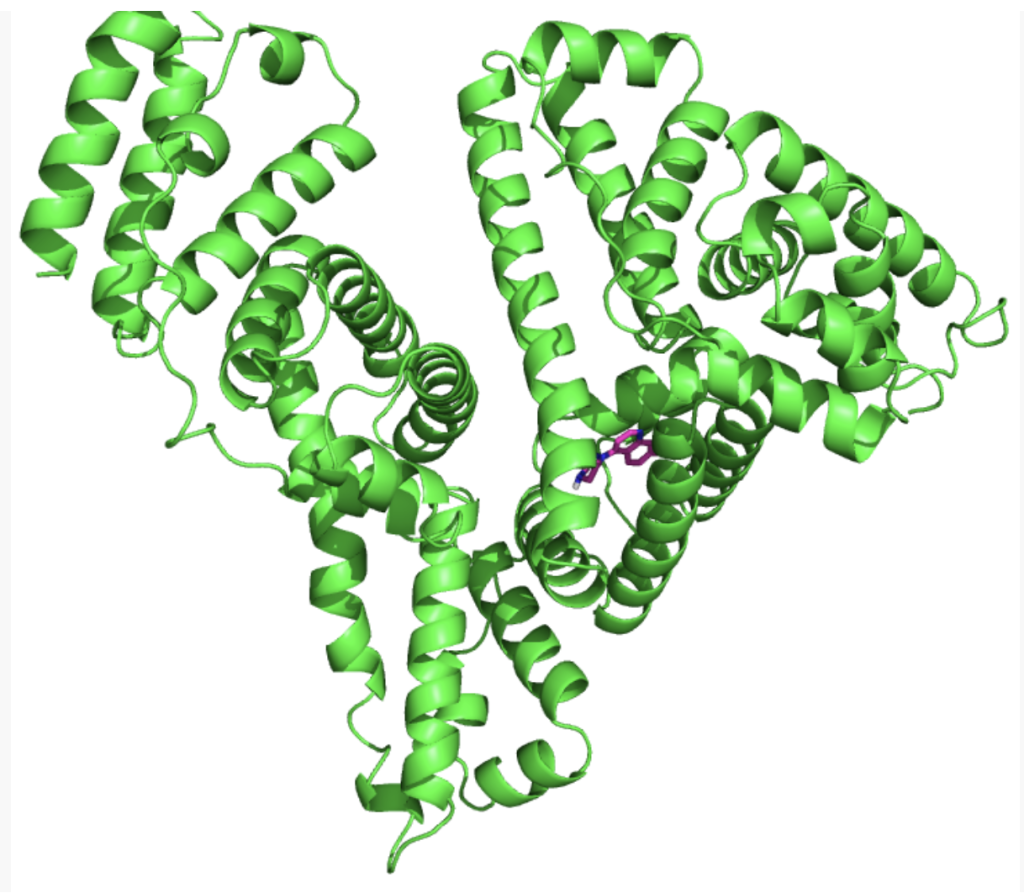
Prediction of 3D structures for proteins, DNA, and RNA provides a foundation for clinical medicine, drug design, and molecular biology research. It not only reveals the structural properties of biological macromolecules but also enables subsequent molecular docking, molecular dynamics, and other studies to assist experimental design and academic publication.
Service Content 2: Drug Molecule Docking Calculation – Drug-Protein, Drug-DNA, Drug-RNA Binding
Molecular docking is based on the “lock-and-key” principle of ligand-receptor interaction, simulating the binding between small molecule ligands and biological macromolecular receptors. It mainly includes protein-small molecule, DNA-small molecule, and RNA-small molecule docking.
Using tools such as AutoDock and Vina, we perform binding energy analysis between drug molecules and target proteins, evaluate binding affinity, conduct structure-activity relationship analysis, predict binding sites, and assist in designing protein structure mutations and drug molecule modifications.
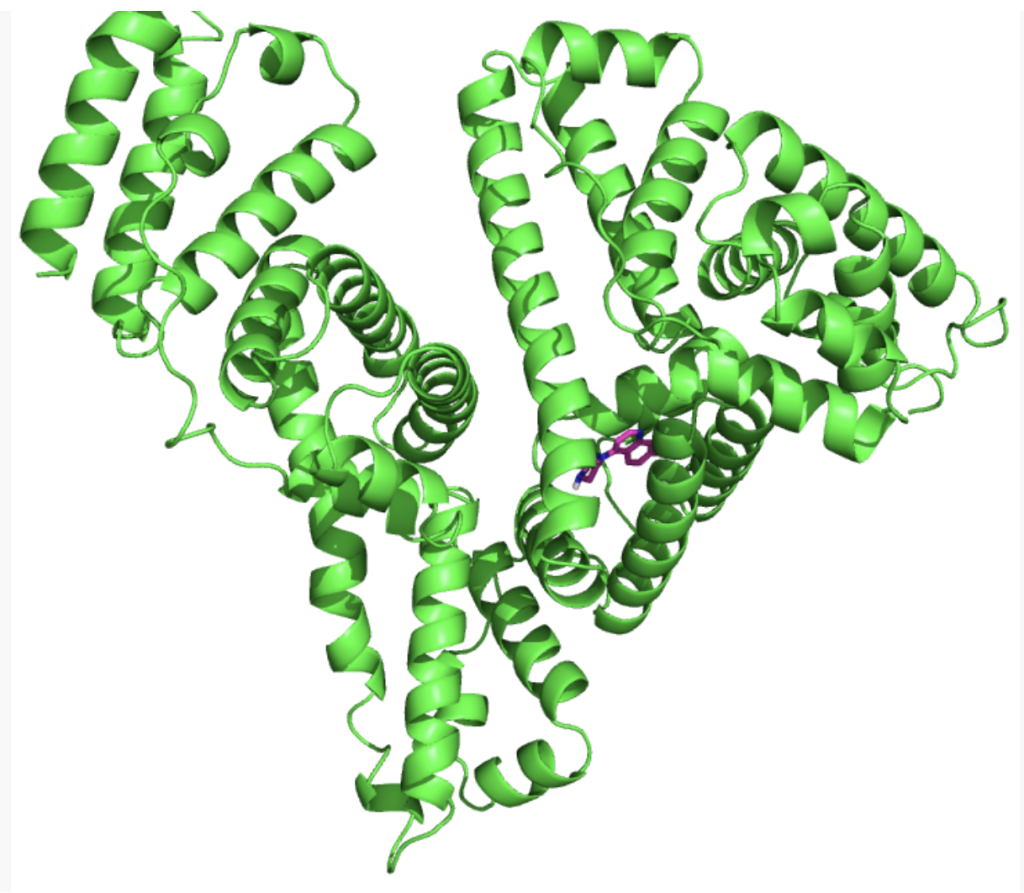
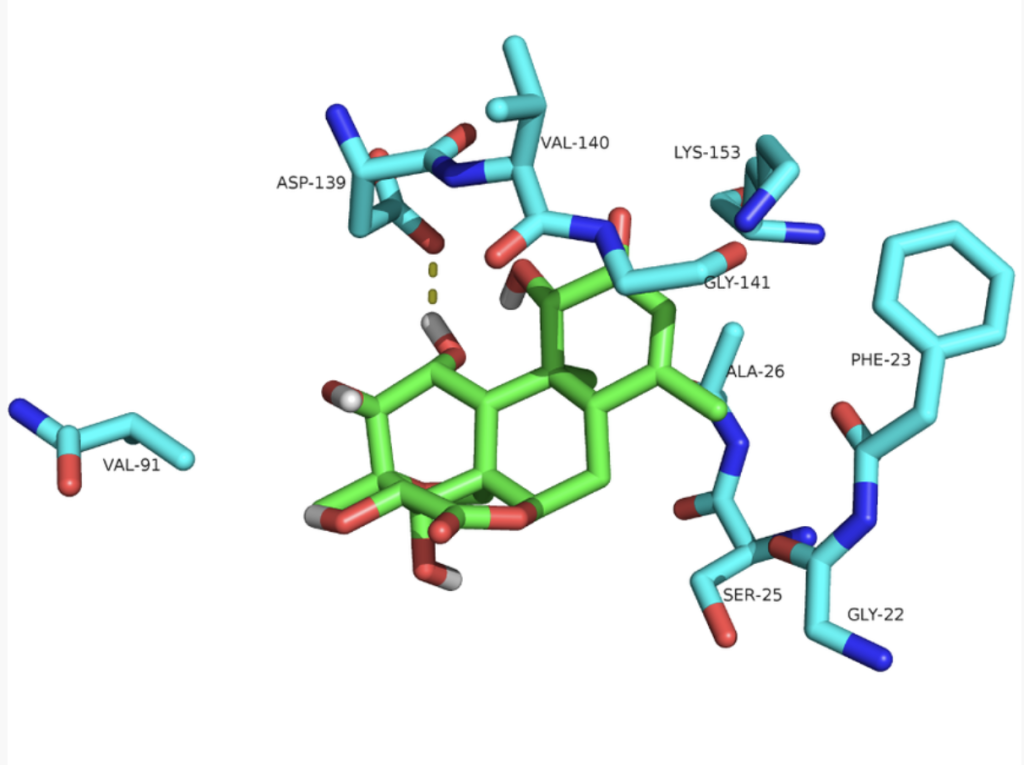
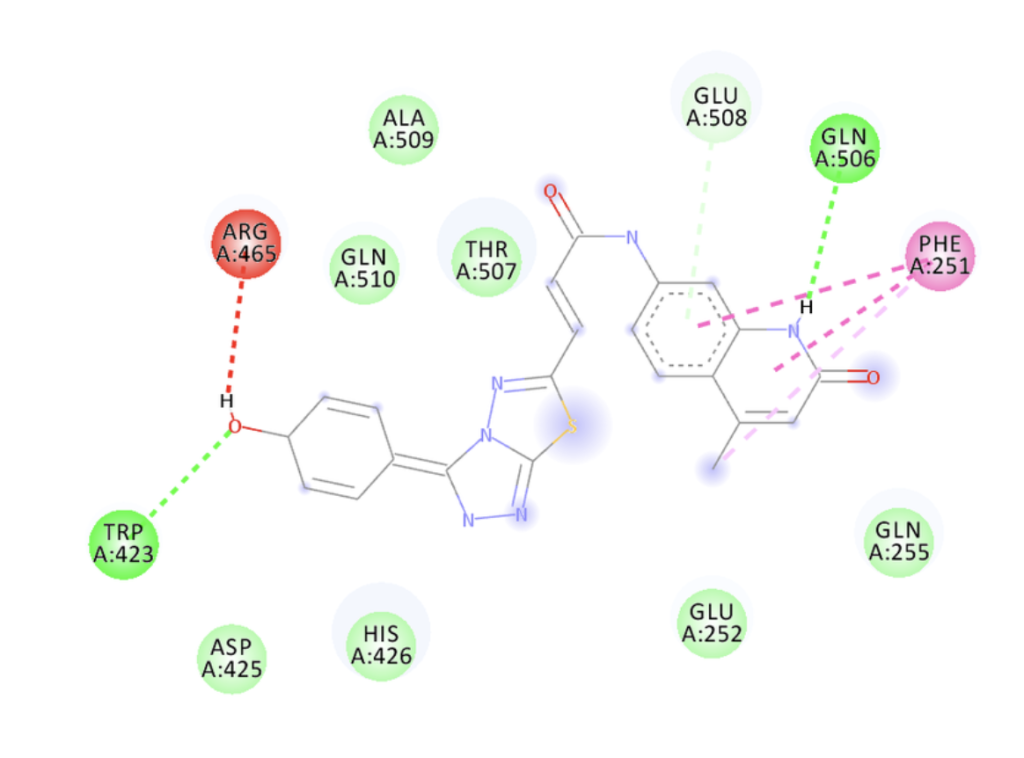
Service Content 3: Macromolecular Docking – Protein-Protein, Protein-DNA, Protein-RNA Binding Prediction
We use HEX, ZDOCK, Rosetta, and other tools to perform protein-protein, protein-DNA, and other macromolecular docking. This verifies results from immunoprecipitation and GST-pull down experiments, rationally infers binding sites, evaluates binding stability and residual energy contributions, and provides guidance for subsequent structural modification and academic writing.
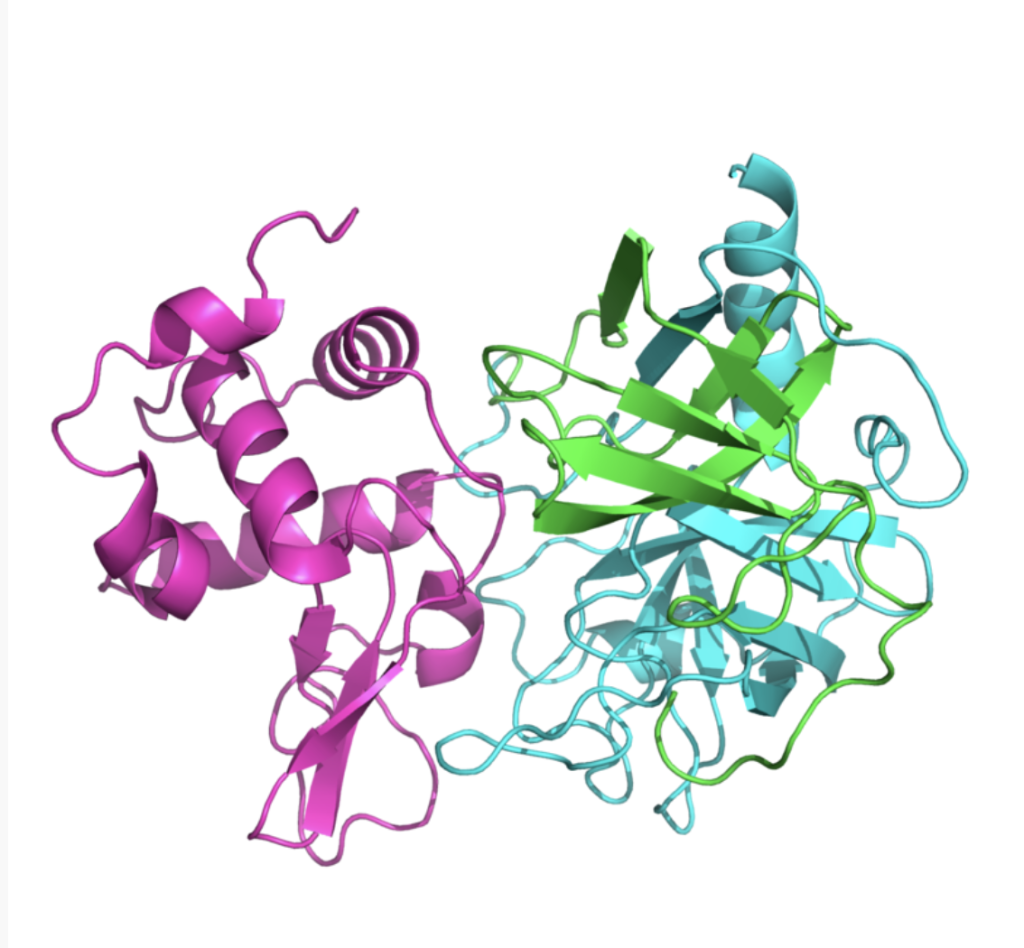
Service Content 4: Molecular Dynamics Simulation – Binding Mode Analysis, Residue Contribution Evaluation, Free Energy Calculation
Through molecular dynamics simulations, we model the dynamic behavior of proteins, DNA, and RNA under varying conditions (temperature, pH, voltage), predict conformational changes, and analyze intermolecular interactions and binding free energy.
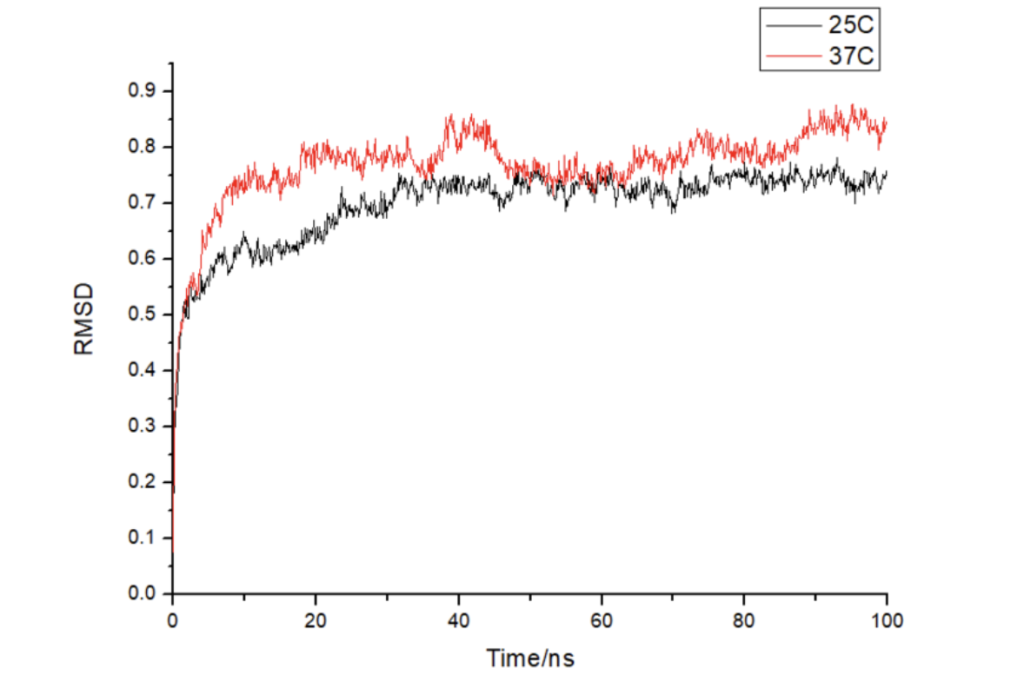
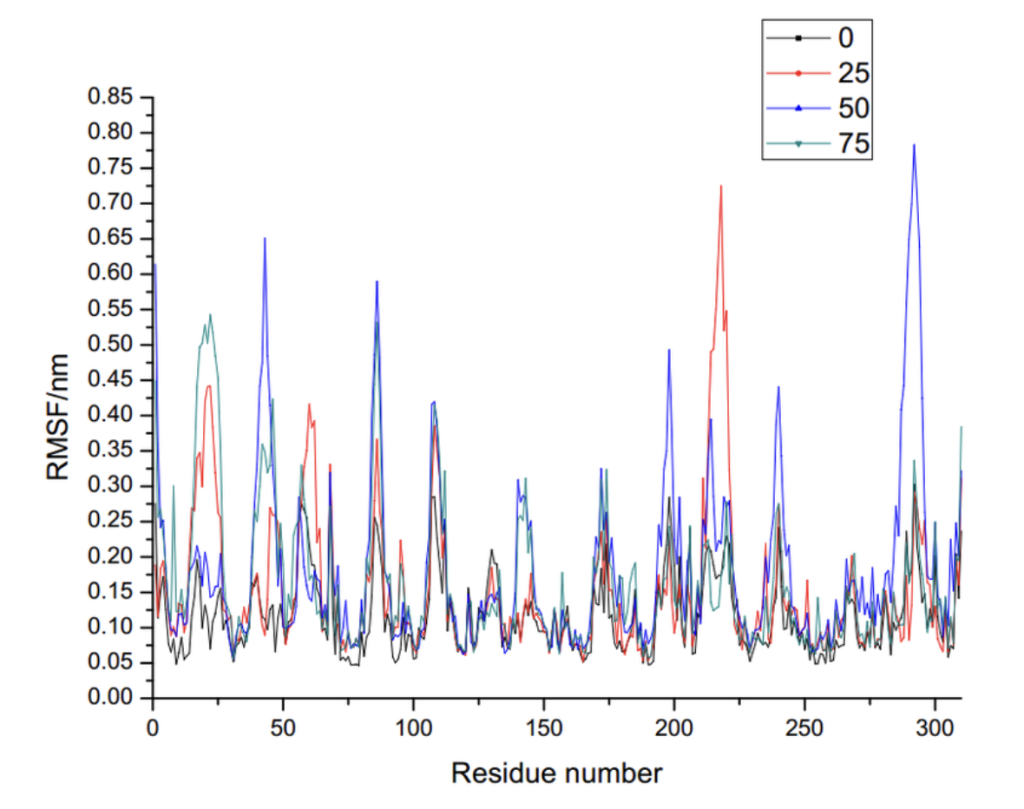
Molecular dynamics can reveal the influence of molecular interactions, evaluate the stability of target structures under different physiological conditions, analyze protein-ligand binding free energy, and guide preclinical and translational research.
Service Content 5: Computer-Aided Drug Screening and Design
Using the Vina simulation software in conjunction with compound databases such as ZINC, we perform large-scale computational virtual screening to identify potential compounds with high binding affinity from the database. We are committed to providing outstanding computer-aided drug design support and services, shortening the cycle of new drug research and development, and offering design solutions for various drug targets.
Combined with protein three-dimensional structure simulation and prediction, we conduct an in-depth analysis of the physicochemical properties around the protein binding pocket, as well as the interaction patterns including hydrophobic and hydrophilic interactions, hydrogen bonding, electrostatic interactions, and van der Waals forces between small molecules and the binding pocket.
Service Content 6: Network Pharmacology and TCM Research
Network pharmacology is a key tool for TCM research. It identifies active ingredients from TCM prescriptions, predicts their potential targets, constructs component-target-pathway networks, and reveals the molecular mechanisms of TCM. This supports TCM modernization and new drug discovery from natural products.
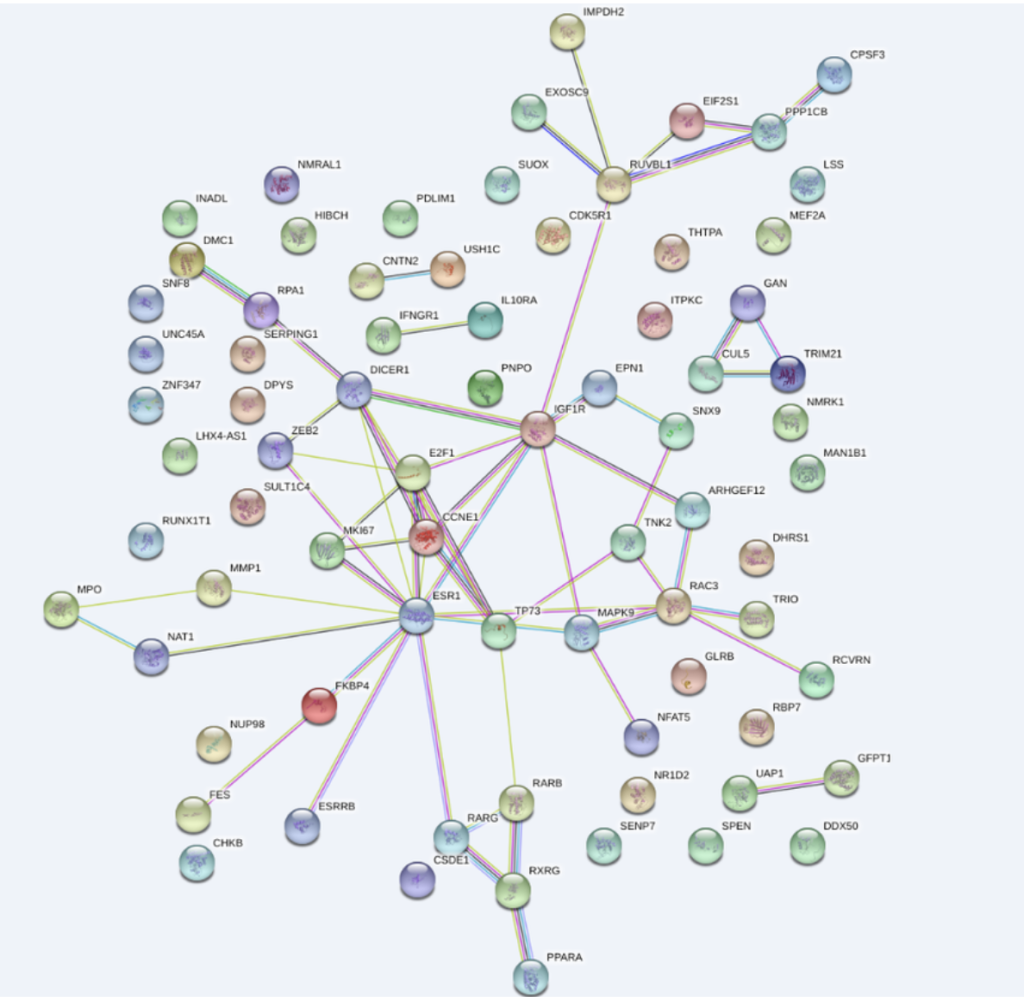
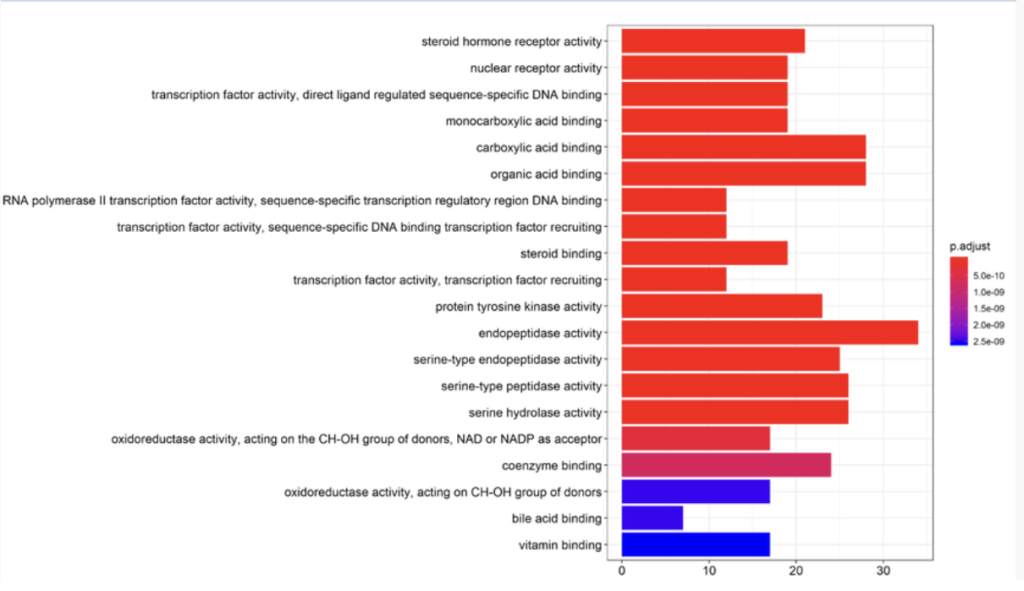
Other Computational Services
- Protein/DNA/RNA 3D structure prediction
- Drug-target, drug-DNA, drug-RNA binding mode analysis
- Protein-protein, protein-DNA, protein-RNA interaction analysis
- Molecular dynamics simulation of proteins, nucleic acids, and complexes
- Residual energy contribution and binding free energy analysis
- Reverse pharmacophore screening to identify potential targets for bioactive molecules
- Network pharmacology analysis
Target Clients
Pharmaceutical R&D, CRO, TCM, organic chemistry, clinical medicine, natural products, biochemistry, molecular biology, food and environmental science, and other related academic and industrial researchers.